GP Regression with a Spectral Mixture Kernel¶
Introduction¶
This example shows how to use a SpectralMixtureKernel module on an ExactGP model. This module is designed for
When you want to use exact inference (e.g. for regression)
When you want to use a more sophisticated kernel than RBF
The Spectral Mixture (SM) kernel was invented and discussed in Wilson et al., 2013.
[1]:
import math
import torch
import gpytorch
from matplotlib import pyplot as plt
%matplotlib inline
%load_ext autoreload
%autoreload 2
In the next cell, we set up the training data for this example. We’ll be using 15 regularly spaced points on [0,1] which we evaluate the function on and add Gaussian noise to get the training labels.
[2]:
train_x = torch.linspace(0, 1, 15)
train_y = torch.sin(train_x * (2 * math.pi))
Set up the model¶
The model should be very similar to the ExactGP model in the simple regression example.
The only difference is here, we’re using a more complex kernel (the SpectralMixtureKernel). This kernel requires careful initialization to work properly. To that end, in the model __init__ function, we call
self.covar_module = gpytorch.kernels.SpectralMixtureKernel(n_mixtures=4)
self.covar_module.initialize_from_data(train_x, train_y)
This ensures that, when we perform optimization to learn kernel hyperparameters, we will be starting from a reasonable initialization.
[5]:
class SpectralMixtureGPModel(gpytorch.models.ExactGP):
def __init__(self, train_x, train_y, likelihood):
super(SpectralMixtureGPModel, self).__init__(train_x, train_y, likelihood)
self.mean_module = gpytorch.means.ConstantMean()
self.covar_module = gpytorch.kernels.SpectralMixtureKernel(num_mixtures=4)
self.covar_module.initialize_from_data(train_x, train_y)
def forward(self,x):
mean_x = self.mean_module(x)
covar_x = self.covar_module(x)
return gpytorch.distributions.MultivariateNormal(mean_x, covar_x)
likelihood = gpytorch.likelihoods.GaussianLikelihood()
model = SpectralMixtureGPModel(train_x, train_y, likelihood)
In the next cell, we handle using Type-II MLE to train the hyperparameters of the Gaussian process. The spectral mixture kernel’s hyperparameters start from what was specified in initialize_from_data.
See the simple regression example for more info on this step.
[6]:
# this is for running the notebook in our testing framework
import os
smoke_test = ('CI' in os.environ)
training_iter = 2 if smoke_test else 100
# Find optimal model hyperparameters
model.train()
likelihood.train()
# Use the adam optimizer
optimizer = torch.optim.Adam(model.parameters(), lr=0.1)
# "Loss" for GPs - the marginal log likelihood
mll = gpytorch.mlls.ExactMarginalLogLikelihood(likelihood, model)
for i in range(training_iter):
optimizer.zero_grad()
output = model(train_x)
loss = -mll(output, train_y)
loss.backward()
print('Iter %d/%d - Loss: %.3f' % (i + 1, training_iter, loss.item()))
optimizer.step()
Iter 1/100 - Loss: 1.281
Iter 2/100 - Loss: 1.258
Iter 3/100 - Loss: 1.232
Iter 4/100 - Loss: 1.212
Iter 5/100 - Loss: 1.192
Iter 6/100 - Loss: 1.172
Iter 7/100 - Loss: 1.156
Iter 8/100 - Loss: 1.143
Iter 9/100 - Loss: 1.131
Iter 10/100 - Loss: 1.121
Iter 11/100 - Loss: 1.114
Iter 12/100 - Loss: 1.110
Iter 13/100 - Loss: 1.106
Iter 14/100 - Loss: 1.103
Iter 15/100 - Loss: 1.101
Iter 16/100 - Loss: 1.098
Iter 17/100 - Loss: 1.094
Iter 18/100 - Loss: 1.089
Iter 19/100 - Loss: 1.085
Iter 20/100 - Loss: 1.080
Iter 21/100 - Loss: 1.076
Iter 22/100 - Loss: 1.072
Iter 23/100 - Loss: 1.069
Iter 24/100 - Loss: 1.067
Iter 25/100 - Loss: 1.065
Iter 26/100 - Loss: 1.063
Iter 27/100 - Loss: 1.061
Iter 28/100 - Loss: 1.060
Iter 29/100 - Loss: 1.057
Iter 30/100 - Loss: 1.054
Iter 31/100 - Loss: 1.051
Iter 32/100 - Loss: 1.048
Iter 33/100 - Loss: 1.044
Iter 34/100 - Loss: 1.039
Iter 35/100 - Loss: 1.035
Iter 36/100 - Loss: 1.029
Iter 37/100 - Loss: 1.023
Iter 38/100 - Loss: 1.015
Iter 39/100 - Loss: 1.006
Iter 40/100 - Loss: 0.995
Iter 41/100 - Loss: 0.981
Iter 42/100 - Loss: 0.965
Iter 43/100 - Loss: 0.946
Iter 44/100 - Loss: 0.924
Iter 45/100 - Loss: 0.898
Iter 46/100 - Loss: 0.870
Iter 47/100 - Loss: 0.839
Iter 48/100 - Loss: 0.806
Iter 49/100 - Loss: 0.770
Iter 50/100 - Loss: 0.731
Iter 51/100 - Loss: 0.686
Iter 52/100 - Loss: 0.637
Iter 53/100 - Loss: 0.583
Iter 54/100 - Loss: 0.523
Iter 55/100 - Loss: 0.460
Iter 56/100 - Loss: 0.394
Iter 57/100 - Loss: 0.327
Iter 58/100 - Loss: 0.260
Iter 59/100 - Loss: 0.194
Iter 60/100 - Loss: 0.133
Iter 61/100 - Loss: 0.078
Iter 62/100 - Loss: 0.032
Iter 63/100 - Loss: -0.005
Iter 64/100 - Loss: -0.040
Iter 65/100 - Loss: -0.086
Iter 66/100 - Loss: -0.144
Iter 67/100 - Loss: -0.206
Iter 68/100 - Loss: -0.264
Iter 69/100 - Loss: -0.313
Iter 70/100 - Loss: -0.354
Iter 71/100 - Loss: -0.392
Iter 72/100 - Loss: -0.430
Iter 73/100 - Loss: -0.472
Iter 74/100 - Loss: -0.518
Iter 75/100 - Loss: -0.567
Iter 76/100 - Loss: -0.616
Iter 77/100 - Loss: -0.662
Iter 78/100 - Loss: -0.703
Iter 79/100 - Loss: -0.740
Iter 80/100 - Loss: -0.775
Iter 81/100 - Loss: -0.816
Iter 82/100 - Loss: -0.860
Iter 83/100 - Loss: -0.902
Iter 84/100 - Loss: -0.940
Iter 85/100 - Loss: -0.975
Iter 86/100 - Loss: -1.008
Iter 87/100 - Loss: -1.042
Iter 88/100 - Loss: -1.077
Iter 89/100 - Loss: -1.112
Iter 90/100 - Loss: -1.145
Iter 91/100 - Loss: -1.173
Iter 92/100 - Loss: -1.199
Iter 93/100 - Loss: -1.226
Iter 94/100 - Loss: -1.254
Iter 95/100 - Loss: -1.279
Iter 96/100 - Loss: -1.302
Iter 97/100 - Loss: -1.321
Iter 98/100 - Loss: -1.341
Iter 99/100 - Loss: -1.361
Iter 100/100 - Loss: -1.379
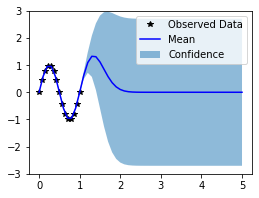
Now that we’ve learned good hyperparameters, it’s time to use our model to make predictions. The spectral mixture kernel is especially good at extrapolation. To that end, we’ll see how well the model extrapolates past the interval [0, 1].
In the next cell, we plot the mean and confidence region of the Gaussian process model. The confidence_region method is a helper method that returns 2 standard deviations above and below the mean.
[7]:
# Test points every 0.1 between 0 and 5
test_x = torch.linspace(0, 5, 51)
# Get into evaluation (predictive posterior) mode
model.eval()
likelihood.eval()
# The gpytorch.settings.fast_pred_var flag activates LOVE (for fast variances)
# See https://arxiv.org/abs/1803.06058
with torch.no_grad(), gpytorch.settings.fast_pred_var():
# Make predictions
observed_pred = likelihood(model(test_x))
# Initialize plot
f, ax = plt.subplots(1, 1, figsize=(4, 3))
# Get upper and lower confidence bounds
lower, upper = observed_pred.confidence_region()
# Plot training data as black stars
ax.plot(train_x.numpy(), train_y.numpy(), 'k*')
# Plot predictive means as blue line
ax.plot(test_x.numpy(), observed_pred.mean.numpy(), 'b')
# Shade between the lower and upper confidence bounds
ax.fill_between(test_x.numpy(), lower.numpy(), upper.numpy(), alpha=0.5)
ax.set_ylim([-3, 3])
ax.legend(['Observed Data', 'Mean', 'Confidence'])